Two requirements are needed for 3D structure reconstruction from projections. One is
projection temp/zeyun (or in our case, particle averages) and the other is projection
angle for each projection (see Fig. 1). With these two inputs, we can reconstruct the 3D
structures using techniques as seen in medical image reconstruction, which will be briefly
described in the following.
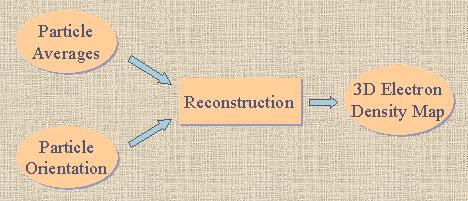 |
| Fig. 1 Pipeline of reconstruction, given particle averages
and their corresponding orientations. |
Particle averages have already been discussed in Classification
. The determination of projection orientation can be accomplished using common line
method or reference-based method. The first method is to determine the relative
orientation between two projections by detecting their common line. The second method assumes
an initial 3D model available and each of the particle temp/zeyun is cross-correlated with each
of the projections of the 3D model. For each particle image, the projection corresponding to the
highest cross-correlation coefficient is assumed to have the same orientation as that particle
image. Clearly the second method can be made iterative to improve the results and can also be
used to solve the particle classification. However, the initial 3D model must be carefully
chosen to guarantee the convergence of this iterative process.
CT/MRI Reconstruction
The CT/MRI reconstruction is based on a well-known theorem called Fourier Slice Theorem,
which says that the Fourier transform of any projection of the original structure must be equal to
a certain zero-crossing slice of the Fourier transform of the original structure. Fig.2(a) shows an
example of a 2D structure reconstruction from its 1D projections. With enough number of projections,
we can compute their Fourier transforms, each of which is "embedded" in the Fourier space and then
an interpolation technique is used to "reconstruct" the Fourier transform of the original
structure f(x, y) in the entire Fourier space (see fig.2 (c)). After we obtain F(u, v), the inverse
Fourier transform of F(u, v) gives the original structure f(x,y). Fig.2 (b) illustrates the overall
pipeline of reconstruction. It is straightforward to extend this procedure into 3D structure
reconstruction.
 |
 |
 |
| (a) Different projections |
(b) Fourier slices and interpolation |
(c) Illustration of reconstruction |
| Fig. 2 2D example of reconstruction pipeline
|
In this section we show one example of reconstruction on P97 dataset (from Dr. Bridget Carragher at The Scripps Research Institute). We used EMAN tool (see [1,2,3]) developed at Baylor College of Medicine, Houston to reconstruct the 3D structures on the above mentioned datasets. However, instead of using the manual or semi-automatic techniques seen in EMNA for picking particles, we used our automatic particle picking algorithm and took the detected particles as input to EMAN.
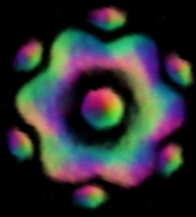
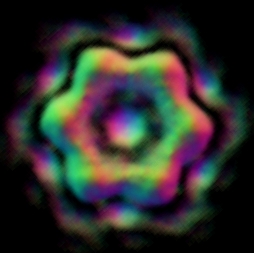
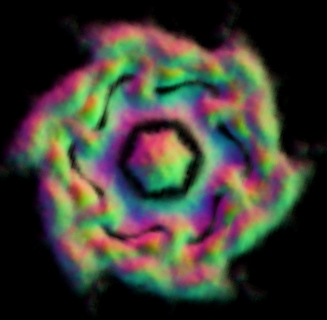

 Fig. 3 Reconstruction results on P97 dataset using EMAN based on the particles boxed out by our algorithm.
Fig. 3 Reconstruction results on P97 dataset using EMAN based on the particles boxed out by our algorithm.
|